
Dr. Alessia Ruggieri
alessia.ruggieri
(at)med.uni-heidelberg.de
Phone: +49 (0)6221 56–7761
Fax: +49 6221 56–4570
Translational Control by RNA Viruses
Projects
Throughout the course of infection, viruses elicit multiple host cell responses, including innate immune response and integrated stress response. The accumulation of viral double-stranded (ds) RNA in the cytosol of infected cells triggers the activation of the stress sentinel Protein Kinase R (PKR), which initiates the integrated stress response by phosphorylating the eukaryotic translation initiation factor eIF2α. This results in the rapid suppression of protein synthesis. The assembly of stress granules (SGs) is frequently observed upon viral infections. SGs are biomolecular condensates that form when polysomes disassemble and stalled mRNAs phase separate from the remaining cytosol together with numerous RNA-binding proteins. The diversity of strategies evolved by viruses to control the translation machinery, overcome translational attenuation and interfere with SGs highlights the importance of the integrated stress response in the host antiviral defense. We aim at uncovering how RNA viruses control or evade cellular responses to ensure their progeny production. To do so, we like to combine integrative and interdisciplinary approaches to study both sides of the coin: the virus and the host. We currently explore the strategies developed by different members of the Flaviviridae family, in particular dengue virus (DENV), how they antagonize or inversely utilize the host stress response and translation machinery. We also tackle the mysteries of the integrated stress response, the fine-tuned regulation of its dynamics, and how it interacts with the innate immune response. Finally, we established a novel microfluidics-based approach to explore the properties of SG phase separation.
Reasearch Interests:
Host Stress Response to RNA Viruses
- Stress Granules and Innate Immune Response
- Translational Repression by Dengue Virus
- RNA Modifications of Flavivirus RNA Genomes
1 | Dynamic Stress Response to RNA Virus Infections
Using long-term live-cell imaging microscopy, we showed that chronic infection with hepatitis C virus in combination with type I interferon (IFN) treatment induces a dynamic host cell stress response that can be visualized by recurring cycles of assembly and disassembly of SGs (Ruggieri et al., 2012). HCV-induced SG-on and SG-off phases is regulated at the level of the eukaryotic initiation factor 2 alpha (eIF2α) by the antagonistic action of the two main switches, PKR, the stress kinase detecting viral double-stranded (ds) RNA, and GADD34, the stress-induced regulatory subunit of Protein Phosphatase 1 (Figure 1) . We recently elucidated the complex regulation of this response with help of quantitative mathematical modelling and discovered that this dynamic stress response is regulated by a stochastic process with memory. Thus, the moment at which cells form SGs is negatively correlated with the duration of the preceding SG phase. This approach also evidenced the importance of inherent cell-to-cell variations in modulating SG dynamics, specifically in the levels of PKR and GADD34 (Klein, Kallenberger et al., 2022). Building on these results, we now question how the establishment of the antiviral response modulates SG formation and cell adaptation to stress during acute viral infection.
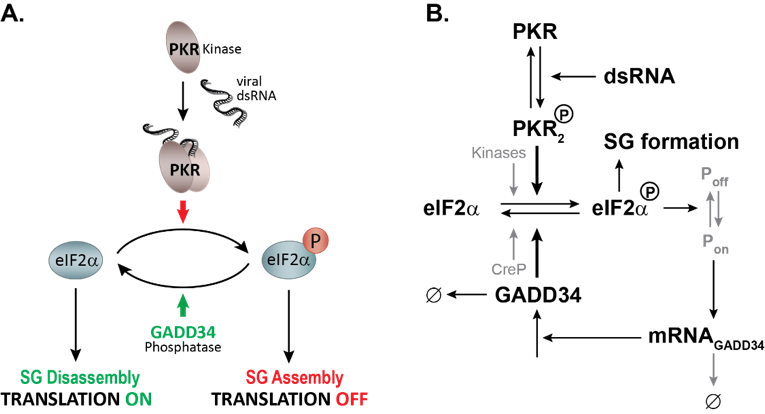
Figure 1: Mathematical modeling of HCV-induced SG oscillations. A. Description of the stress signaling pathway activated by HCV infection. PKR is activated by binding to viral dsRNA, dimerizes and autophosphorylates. Active PKR phosphorylates its direct substrate eIF2α leading to a bulk translation initiation shutoff, polysome disassembly and assembly of SGs. To promote survival, cells GADD34 is transcriptionally and translationally upregulated and in complex with the Protein Phosphatase 1 dephosphorylates eIF2α. As translation is resumed, SGs will disassemble. As long as viral dsRNA is present in the cell, these cycles of active and stalled translation will occur. B. Simplified schematics of the mathematical model of the integrated stress response to dsRNA.
2 | Regulation of GADD34, the Stress-induced Regulatory Subunit of Phosphatase PP1
A key prediction of our mathematical model of the integrated stress response was the key role of GADD34, both at the protein and at the mRNA level, in the adaptation to repeated stress. We recently identified an AU-rich element (ARE), a regulatory element known to destabilize mRNAs, in the 3′ untranslated region (UTR) of PPP1R15A mRNA, encoding GADD34 (Figure 2). Under normal conditions, we found that PPP1R15A ARE is recognized by proteins of the TTP family that promote rapid mRNA decay. Upon exposure to different types of stress, as TTP proteins are inactivated, PPP1R15A mRNA is transiently stabilized, allowing for rapid GADD34 translation and promoting entry into the stress adaptation phase. With this work, we uncovered the role of PPP1R15A mRNA turnover in shaping the dynamics of stress adaptation and cell sensitivity to repetitive stress exposure and identified GADD34 as the molecular memory of the activated integrated stress response (Magg, Manetto et al., 2024). By studying the post-transcriptional regulation of PPP1R15A mRNA, we have realized that basal levels of GADD34 are critical and regulated by additional mechanisms, which we are currently exploring.
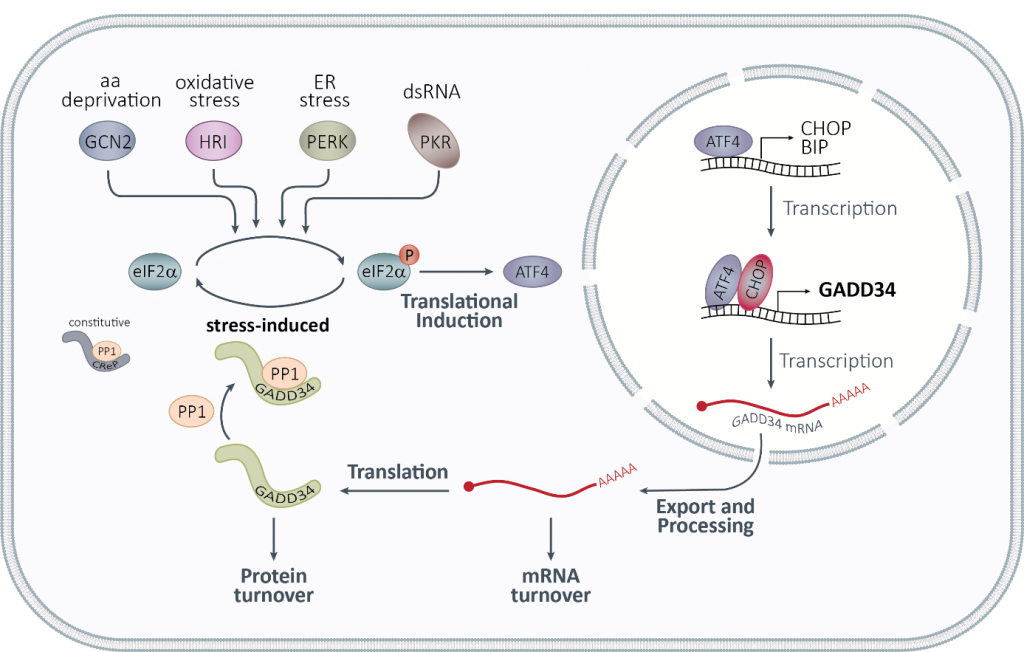
Figure 2: The integrated stress response.
3 | Assembly of Stress Granule like Condensates in Droplets
Bottom up approaches making use of purified SG proteins such as G3BP1, the main SG nucleating protein, have revealed exciting features of how these biomolecular condensates dynamically form by the phase separation. However, in cells, SG condensation also results from multiple interactions with RNAs and RNA-binding proteins, which greatly complexify this model. To tackle this challenge, we opted for the combined use of simple microfluidics and fluorescence microscopy, to establish a droplet-based method that bridges the gap between in vitro systems and living cells, providing a platform to study stress-induced phase separation under physiological conditions.
4 | Translational Control by Flaviviruses
Flaviviruses such as DENV, ZIKV and WNV have a positive sense single-stranded RNA genome with a type I cap at their 5’ end and a non-polyadenylated 3′ untranslated region. Viral genomes therefore compete with host mRNAs for their translation. We have shown that flavivirus infection induces a severe repression of the host cell translation in human cells, which is uncoupled from the cellular stress response (Roth et al., 2017). Despite this repression, translation of viral genomes is maintained while host translation is repressed. This work suggested an unconventional and virus-specialized translation initiation mechanism that we are currently investigating.
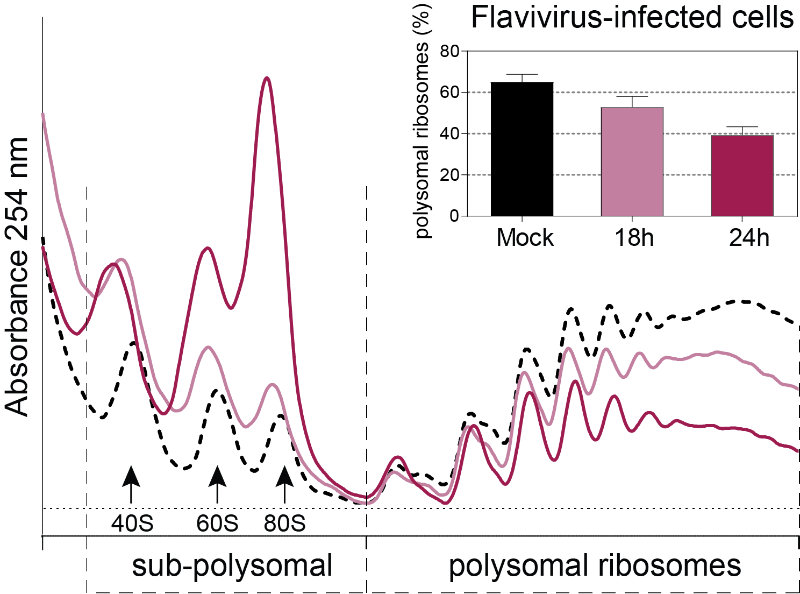
Figure 3: Polysome profiles of Flavivirus-infected Huh7 cells. This technique allows the identification of translational changes in host cells submitted to different environmental stresses including virus infection by separating heavier ribosome-associated mRNAs (actively translating mRNAs, polysomal ribosomes) from the light sub-polysomal mRNAs (Poorly or not translated mRNAs, sub-polysomal ribosomes). Flaviviruses reduce the bulk of active translating mRNAs in the course of the infection.
5 | Role of RNA Modifications in the Flavivirus Life Cycle
Chemical modifications of RNA affect all stages of the RNA life cycle, including splicing, stability and translation. RNA modifications are also essential for the host immune system to distinguish between self and foreign RNA. Increasing evidence suggests that viral RNA genomes, whose structure closely mimics that of cellular mRNAs, are also highly modified. However, results on their detection and mapping are partly controversial. We have decided to take on this problem by developing new purification approaches and we aim to explore the epitranscriptomics of viral genome RNA over the course of flavivirus infection. This will enable us to assess its impact on the virus life cycle and host recognition.
Funding



Selected Publications
Link to complete publication list: ORCID-iD: 0000–0001-9981–3308
- Wu CC, Naarmann-de Vries IS, Hartmann J, Nidoieva Z, Kopietz K, Marchand V, Özrendeci Z, Lindner D, Schelchshorn S, Flad S, Frye M, Papavasiliou N, Schirmeister T, Stoecklin G, Schott J, Motorin Y, Tuorto Y, Dieterich C, Helm M, Ruggieri A. The two-step purification method ViREn identifies a single NSUN6-mediated 5‑methylcytosine modification promoting dengue virus RNA genome turnover. 2025. BioRxiv. doi: 10.1101/2025.03.17.643699.
- Magg V*, Manetto A*, Kopp K, Wu C, Lindner D, Naghizadeh M, Schott J, Eke L, Welsch J, Kallenberger SM, Haucke V, Locker N, Stoecklin G†, Ruggieri A†. Turnover of PPP1R15A mRNA encoding GADD34 controls responsiveness and adaptation to cellular stress. 2024. Cell Reports 43(4):114069.
- Klein P*, Kallenberger SM*, Roth H, Roth K, Ly-Hartig TBN, Magg V, Aleš J, Talemi SR , Qiang Y, Wolf S, Oleksiuk O, Kurilov R, Di Ventura B, Bartenschlager R, Eils R, Rohr K, Hamprecht FA, Höfer T, Fackler OT, Stoecklin G and Ruggieri A. Temporal control of the integrated stress response by a stochastic molecular switch. 2022. Science Advances 8(12):eabk2022.
- Talemi SR, Bartenschlager M, Schmid B, Ruggieri A†, Bartenschlager R†, Höfer T†. Dengue virus is sensitive to inhibition prior to productive replication. 2021. Cell Reports 37(2):109801.
- Ruggieri A, Helm M, Chatel-Chaix L. An epigenetic “extreme makeover”: the methylation of flaviviral RNA (and beyond). 2020. RNA Biol 18:1–13.
- Eiermann N, Haneke K, Sun Z, Stoecklin G, Ruggieri A. Dance with the devil: Stress granules and signaling in antiviral responses. 2020. Viruses 12(9), 984.
- Haneke K, Schott J, Lindner D, Hollensen AK, Damgaard CK, Mongis C, Knop M, Palm W, Ruggieri A, Stoecklin G. CDK1 couples proliferation with protein synthesis. 2020. J Cell Biol. 219(3): e201906147.
- Brocard M, Iadevaia V, Klein P, Hall B, Lewis G, Lu J, Burke J, Willcocks M, Parker R, Goodfellow IG, Ruggieri A, Locker N. Norovirus infection results in eIF2α-independent host translation shut-off and remodels the G3BP1 interactome evading stress granule formation. 2020. PLoS Pathog. 16(1):e1008250.
- Ruggieri, A. and Stoecklin, G. (2019). A Signal to Condense. Nat Chem Biol. 15(1):5–6.
- Roth, H., Magg, V., Uch, F., Mutz, P., Klein, P., Haneke, K., Lohmann, V., Bartenschlager, R., Fackler, O.T., Locker, N., Stoecklin, G., Ruggieri, A. (2017). Flavivirus infection uncouples translation suppression from cellular stress responses. mBio 8(1):e02150-16.
- Ruggieri, A., Dazert, E., Metz, P., Hofmann, S., Bergeest, J.-P., Mazur, J., Bankhead, P., Hiet, M.-S., Kallis, S., Alvisi, G., Samuel, C.E., Lohmann, V., Kaderali, L., Rohr, K., Frese, M., Stoecklin, G., Bartenschlager, R. (2012). Dynamic oscillation of translation and stress granule formation mark the cellular response to virus infection. Cell Host Microbe 12(1): 71–85.
- Ruggieri Lab Molecular Virology (Heidelberg University Hospital)
- SFB1129 “Integrative Analysis of Pathogen Replication and Spread” Project TP13
- TRR186 “Molecular Switches in the Spatio-Temporal Control of Cellular Signal Transmission” Project A14
- TRR319 RMaP “RNA Modifications and Processing” Project A05
- TRR319 YouRMaP “Young Researchers in RMaP” Integrated Research Training Group
- The Heidelberg RNA/Translation Club (RNA Society Salon)

